Neues
07.11.2022
Heidelberger Technologieunternehmen EML und alfaview® unter einem Dach
Erweiterung der Videokonferenzsoftware alfaview® um intelligente Spracherkennung und -protokollierung
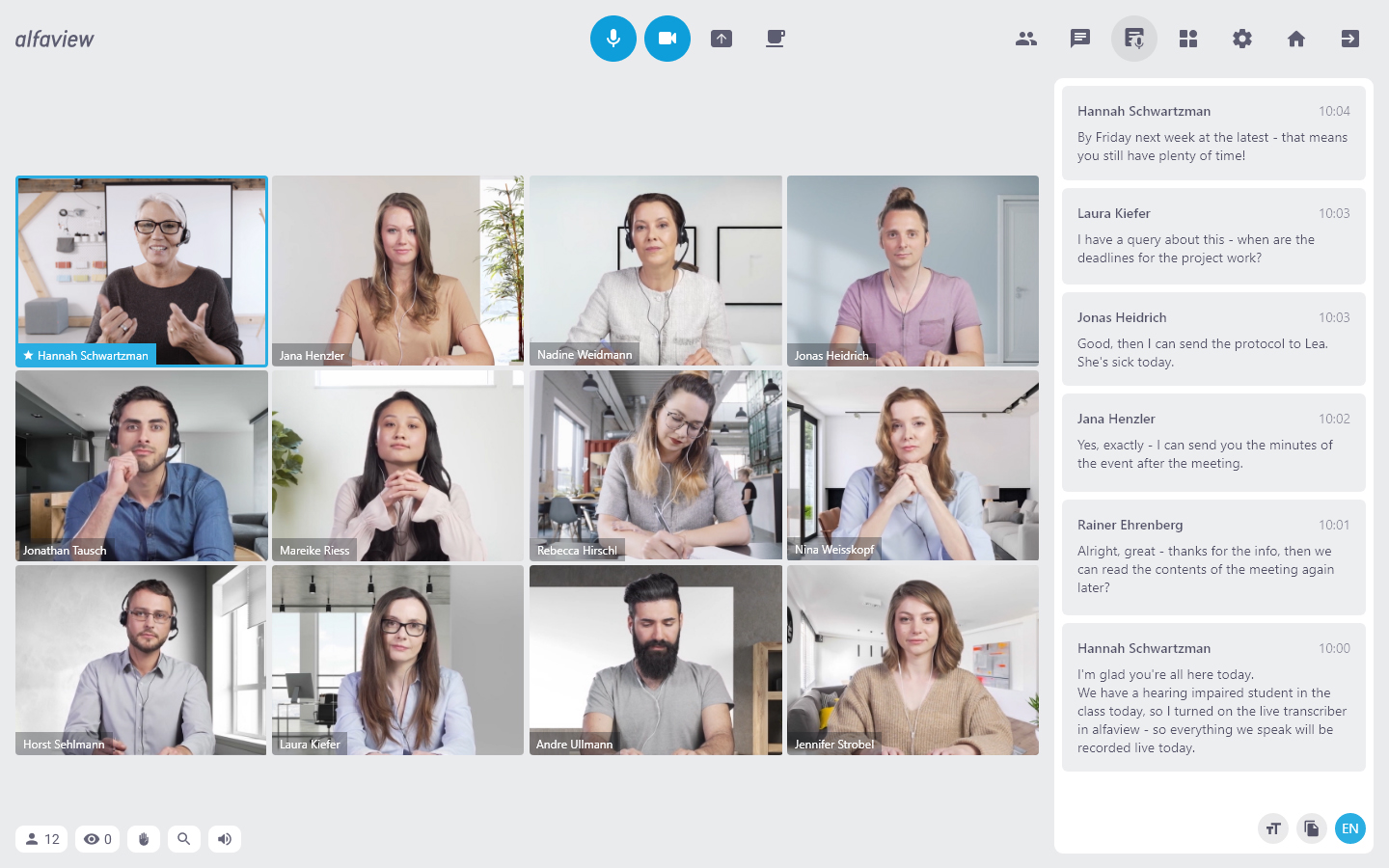
Zur Erweiterung der Videokonferenzsoftware alfaview hat die alfa®-Unternehmensgruppe des Geschäftsführers und Gründers Niko Fostiropoulos im September 2022 das Heidelberger Technologieunternehmen EML Speech Technology GmbH erworben. Mit dem Kauf von EML durch die Muttergesellschaft alfacube firmiert EML nun unter demselben Dach wie die Tochtergesellschaft alfaview.
Die EML Speech Technology GmbH ist ein Entwicklungs- und Forschungsunternehmen auf dem Gebiet der automatischen Spracherkennung, das von dem Physiker und SAP-Mitbegründer Klaus Tschira ins Leben gerufen wurde. EML entwickelt auf Basis neuester Verfahren und Algorithmen im Bereich der künstlichen Intelligenz und des maschinellen Lernens.
Mit dem Erwerb der Heidelberger Forschungs- und Entwicklungsschmiede für digitale Sprachtranskription möchte Niko Fostiropoulos die Videokonferenzsoftware alfaview erweitern und intelligente Protokollierungs- und Transkriptionsinstrumente für virtuelle Meetings zur Verfügung stellen. Mit EML ist alfaview die einzige DSGVO-konforme Videokonferenzplattform mit eigener Sprachtranskription, made in Germany.
Während US-Anbieter auf Tools setzen, die die Daten der europäischen Anwender:innen in die USA oder andere Drittstaaten übertragen, setzt alfaview auf die Erweiterung der eigenen Unternehmensfamilie und schafft mit dem Kauf von EML die Basis für umfangreiche KI-Sprachtechnologien auf Grundlage der Datenschutzgrundverordnung.
EML wird auch weiterhin vom Standort Heidelberg aus mit demselben Team von Sprachwissenschaftler:innen, Mathematiker:innen und Informatiker:innen state-of-the-art Lösungen im Bereich der Sprachtechnologie entwickeln. Über die bereits bestehenden on-premise-Lösungen soll jedoch eine hochskalierbare, dynamische und 100% DSGVO-konforme Cloud für die Sprachdienste von EML entwickelt und am Markt zur Verfügung gestellt werden.
Neben der Implementierung der Sprachtechnologie in das Produkt alfaview haben sich die Unternehmen alfaview, alfatraining und EML mit der Hochschule Karlsruhe (Prof. Dr.-Ing. Matthias Wölfel) sowie der Universität Hohenheim (Prof. Dr. Jens Vogelgesang) auch auf ein zukunftsweisendes BMBF-Forschungsprojekt beworben, in welchem die Partner zeigen möchten, wie durch den Einsatz von virtuellen Räumen, Distanzen überwunden werden und zur Verbesserung der Lebensqualität führen können.
Ziel ist es, die Technologie von EML so weiterzuentwickeln, dass Menschen aus allen Ländern der Welt barrierefrei in einem Meeting kommunizieren können und unterschiedliche Sprachen aufgrund der intelligenten Technik keine Rolle mehr spielen.
Mit dem strategischen Kauf des Technologieexperten EML gewinnt alfaview Videoconferencing Systems ein Produktportfolio rund um das zukunftsweisende Thema Sprachtranskription, das das Knowhow des Konzerns von Niko Fostiropoulos leistungsstark ergänzen wird.
22.09.2020
EML Speech Technology ist Partner im Forschungsprojekt „AdaptAR"

Handbücher, die mehr wissen: Dank Augmented Reality und digitalem Zwilling zu besserer und kostengünstigerer Produktdokumentation
Jeder kennt sie: Anleitungen und Handbücher, die nicht die Erwartungen und Bedürfnisse ihrer Nutzer erfüllen. Entweder fehlen wichtige Informationen oder nachträgliche Änderungen konnten vom Hersteller nicht mehr in der gedruckten Gebrauchsanweisung berücksichtigt werden. Zudem benötigen die Anwender je nach ihrem technischen Wissen unterschiedlich aufbereitete Angaben. Die Folge: Vielen Anwendern sind komplexe Handbücher ein Graus, da sie nicht in der Lage sind, die passenden Antworten auf konkrete Fragen zu einem Produkt zu finden. Mehr Informationen
11.02.2019
EML auf der CCW: Wer spricht denn da?
Die EML European Media Laboratory GmbH zeigt auf der internationalen Messe CCW 2019 in Berlin, wie Sprachtechnologie im Call-Center für mehr Effizienz und Qualitätskontrolle sorgen kann: durch die automatische Verschriftung, die Erkennung verschiedener Sprecher und eine Stichwortsuche in Echtzeit. Das Heidelberger Unternehmen setzt dafür unter anderem Verfahren der Künstlichen Intelligenz ein. Zur Pressemitteilung
24.01.2019
TAPAS - Training in Lissabon
EML ist Partner von "TAPAS", dem "Training Network on Automatic Processing of PAthological Speech." Das Netzwerk mit neun Partnern aus 5 europäischen Ländern will Menschen mit krankheitsbedingten Sprachstörungen, zum Beispiel durch Parkinson oder Schlaganfall, durch den Einsatz von Sprachtechnologie unterstützen. Das nächste Trainingsevent findet vom 11.-14.02.2019 in Lissabon, Portugal, statt. Mehr Informationen
20.06.2018
"Sprachanalyse profitiert von künstlicher Intelligenz"
Markus Klehr, Leiter Entwicklung, und Dr. Volker Fischer, Leiter Forschung, im Gespräch über Transkription, den Einsatz im Call-Center und über europäische Datenschutzrichtlinien. Zum Artikel
11.06.2018
"Ein Qualitätssprung für die Sprachtechnologie"
Dr. Volker Fischer, Leiter Forschung des EML, spricht in der "Technology Review" über den Einfluss maschineller Lernverfahren auf die Spracherkennung. zum Artikel
29.05.2018
Innovativ durch Forschung: EML erhält Gütesiegel des Stifterverbands
Der Stifterverband für die Deutsche Wissenschaft hat das EML mit dem Gütesiegel "Innovativ durch Forschung" ausgezeichnet. DAs Heidelberger Forschungs- und Entwicklungsunternehmen erhält das Gütesiegel bereits zum zweiten Mal hintereinander.
03.04.2018
EML ist Partner des EU-Netzwerks "TAPAS"
EML ist Partner von "TAPAS", dem "Training Network on Automatic Processing of PAthological Speech." Das Netzwerk mit neun Partnern aus 5 europäischen Ländern will Menschen mit krankheitsbedingten Sprachstörungen, zum Beispiel durch Parkinson oder Schlaganfall, durch den Einsatz von Sprachtechnologie unterstützen. Mehr Informationen
01.03.2018
Zur Sprache gebracht: Kommunikation, wie man sie braucht
Intelligenter Anrufbeantworter hilft Makler, immer erreichbar zu bleiben. „Best-Practice"-Beispiele für den Einsatz von EML-Sprachtechnologie (Teil 4) – Messestand und Demonstration live auf der CCW in Berlin vom 27. Februar bis 1. März. Zur Pressemitteilung
28.02.2018
Zur Sprache gebracht: Die kooperative Alternative
Das deutsche Forschungsprojekt „KoFFI" nutzt EML-Sprachtechnologie für teilautomatisiertes Autofahren. „Best-Practice"-Beispiele für den Einsatz von EML-Sprachtechnologie (Teil 3) – Messestand und Demos live auf der CCW in Berlin vom 27. Februar bis 1. März. Zur Pressemitteilung
27.02.2018
Zur Sprache gebracht: Finanzinstitut optimiert Kundenkommunikation mit Sprachanalyse
ASC Technologies, weltweit führender Softwareanbieter im Bereich Multi-Channel-Recording,
Qualitätsmanagement und Analytics, baut auf EML-Sprachtechnologie. „Best-Practice"-Beispiele für den Einsatz von EML-Sprachtechnologie (Teil 2) – Messestand und Demonstration live auf der CCW in Berlin vom 27. Februar bis 1. März. Zur Pressemitteilung
26.02.2018
Zur Sprache gebracht: Und das intelligente Haus „hört zu"
„Das EU-Projekt „LISTEN" („Hör zu") entwickelt ein verlässliches Freisprechsystem für intelligentes Wohnen. „Best-Practice"-Beispiele für den Einsatz von EML-Sprachtechnologie (Teil 1) – Messestand und Demos live auf der CCW in Berlin vom 27. Februar bis 1. März. Zur Pressemitteilung
20.02.2018
EML auf der CCW 2018
Auch auf der CCW 2018 im Februar ist das EML wieder mit einem eigenen Stand vertreten und zeigt auf dieser internationalen Messe zur Kundenkommunikation die Vorzüge von Sprachanalyse in Echtzeit. Ticketgutscheine für einen Besuch am EML-Stand.
18.10.2017
Do you want KoFFI?
Statustagung des BMBF-Projekts "KoFFI" (Kooperative Fahrer-Fahrzeug-Interaktion) beim EML im Mathematikon Heidelberg. Mit den Partnern Bosch, Daimler, Universität Ulm, Hochschule Heilbronn und Hochschule der Medien Stuttgart. Impressionen










